Comparative analysis of conserved miRNAs in maize germination and vegetative development
DOI:
https://doi.org/10.17268/sci.agropecu.2026.037Palavras-chave:
microRNA, maize, seed germination, ribosomal proteins, Northern blotResumo
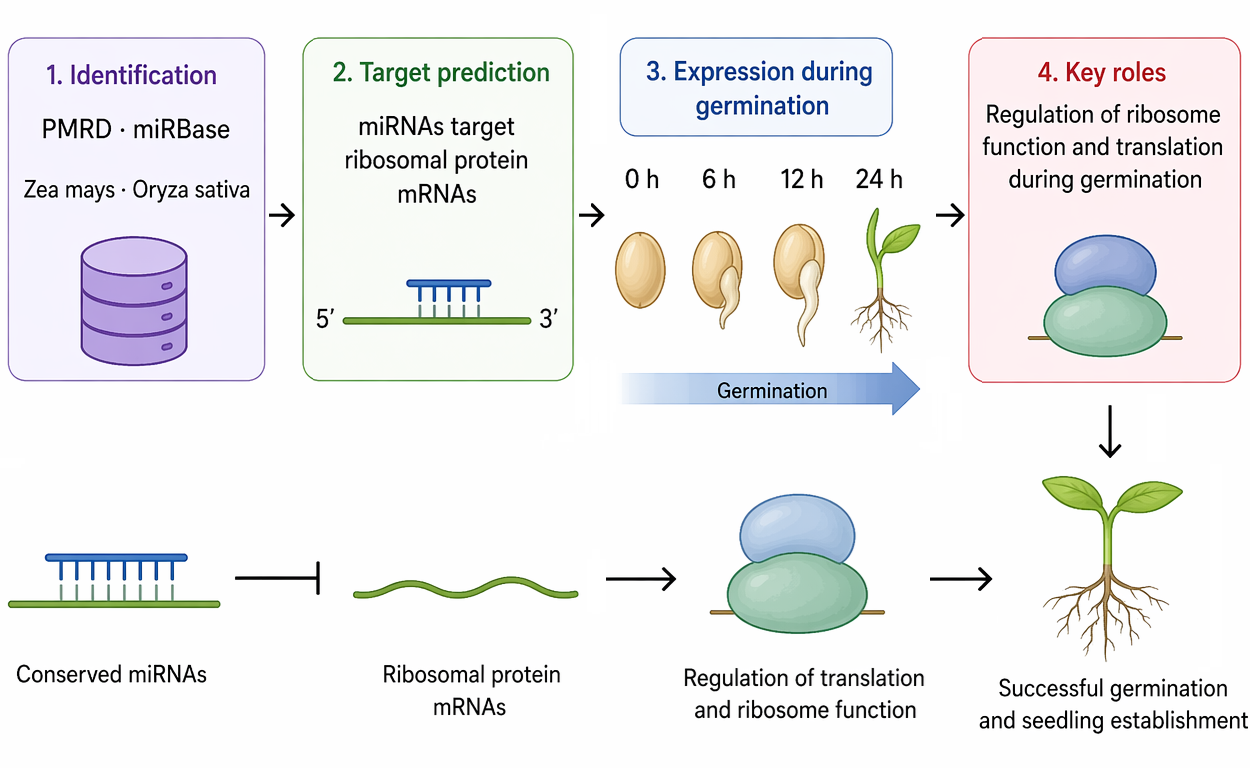
Seed germination represents a metabolically active but transcriptionally limited transition that depends largely on post-transcriptional regulation. MicroRNAs (miRNAs) are key regulators of gene expression through mRNA slicing or translational inhibition, and may play an important role in controlling stored mRNAs during early germination. In this study, we combined bioinformatic analysis and experimental validation to identify conserved miRNAs involved in maize (Zea mays) germination. Mature miRNA sequences from Zea mays and Oryza sativa were retrieved from PMRD and miRBase v22.1, identifying conserved candidates across cereals. From this set, ten miRNAs were selected based on sequence conservation and reported roles in seed biology. Target prediction using thermodynamic modelling (ViennaRNA; MFE ≤ −25 kcal/mol) revealed potential interactions between specific miRNAs and mRNAs encoding ribosomal proteins, suggesting regulation of the translational machinery. Expression analysis by Northern blot across germination stages (0, 6, 12, and 24 h) and vegetative tissues showed distinct patterns. miR528 and miR396a were highly expressed in seeds but absent in vegetative tissues, while miR408 decreased after imbibition. miR414 and miR415 displayed transient expression during germination. Notably, miR160a was strongly expressed during germination, suggesting a role beyond vegetative development. These results support a model in which miRNAs regulate stored mRNAs and translation during germination. This work provides candidate miRNAs for functional studies and potential targets for improving seed vigor and early seedling establishment in maize.
Referências
Bai, B., van der Horst, S., Cordewener, J. H. G., America, T. A. H. P., Hanson, J., & Bentsink, L. (2020). Seed-stored mRNAs that are specifically associated to monosomes are translationally regulated during germination. Plant Physiology, 182(1), 378–392. https://doi.org/10.1104/pp.19.00644
Basbouss-Serhal, I., Pateyron, S., Cochet, F., Leymarie, J., & Bailly, C. (2017). 5' to 3' mRNA decay contributes to the regulation of Arabidopsis seed germination by dormancy. Plant Physiology, 173(3), 1709–1723. https://doi.org/10.1104/pp.16.01933
Cock, P. J. A., Antao, T., Chang, J. T., Chapman, B. A., Cox, C. J., Dalke, A., Friedberg, I., Hamelryck, T., Kauff, F., Wilczynski, B., & de Hoon, M. J. L. (2009). Biopython: Freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics, 25(11), 1422–1423. https://doi.org/10.1093/bioinformatics/btp163
Curaba, J., Singh, M. B., & Bhalla, P. L. (2014). miRNAs in the crosstalk between phytohormone signalling pathways. Journal of Experimental Botany, 65(6), 1425–1438. https://doi.org/10.1093/jxb/eru002
Ding, D., Wang, Y., Han, M., Fu, Z., Li, W., Liu, Z., & et al. (2012). MicroRNA transcriptomic analysis of heterosis during maize seed germination. PLoS ONE, 7(6), e39578. https://doi.org/10.1371/journal.pone.0039578
Ding, N., & Zhang, B. (2023). microRNA production in Arabidopsis. Frontiers in Plant Science, 14, 1096772. https://doi.org/10.3389/fpls.2023.1096772
Donayre-Torres, A. J., Esquivel-Soto, E., Gutiérrez-Xicoténcatl, M. de L., Esquivel-Guadarrama, F. R., & Gómez-Lim, M. A. (2009). Production and purification of immunologically active core protein p24 from HIV-1 fused to ricin toxin B subunit in E. coli. Virology Journal, 6, 17. https://doi.org/10.1186/1743-422X-6-17
Dong, Q., Hu, B., & Zhang, C. (2022). microRNAs and their roles in plant development. Frontiers in Plant Science, 13, 824240. https://doi.org/10.3389/fpls.2022.824240
Gao, Y., Feng, B., Gao, C., Zhang, H., Wen, F., Tao, L., Fu, G., & Xiong, J. (2022). The evolution and functional roles of miR408 and its targets in plants. International Journal of Molecular Sciences, 23(1), 530. https://doi.org/10.3390/ijms23010530
Gran, P., Visscher, T. W., Bai, B., Nijveen, H., Mahboubi, A., Bakermans, L. L., Willems, L. A. J., & Bentsink, L. (2025). Unravelling the dynamics of seed-stored mRNAs during seed priming. New Phytologist, 247(5), 2196–2209. https://doi.org/10.1111/nph.70098
Guo, Z., Kuang, Z., Deng, Y., Li, L., & Yang, X. (2022). Identification of species-specific microRNAs provides insights into dynamic evolution of microRNAs in plants. International Journal of Molecular Sciences, 23(22), 14273. https://doi.org/10.3390/ijms232214273
Gutierrez, L., Mongelard, G., Floková, K., Pacurar, D. I., Novák, O., Staswick, P., Kowalczyk, M., Pacurar, M., Demailly, H., Geiss, G., & Bellini, C. (2012). Auxin controls Arabidopsis adventitious root initiation by regulating jasmonic acid homeostasis. Plant Cell, 24(6), 2515–2527. https://doi.org/10.1105/tpc.112.099119
Hunter, J. D. (2007). Matplotlib: A 2D graphics environment. Computing in Science & Engineering, 9(3), 90–95. https://doi.org/10.1109/MCSE.2007.55
Jiang, A., Guo, Z., Pan, J., Yang, Y., Zhuang, Y., Zuo, D., Hao, C., Gao, Z., Xin, P., Chu, J., Zhong, S., & Li, L. (2021). The PIF1-miR408-PLANTACYANIN repression cascade regulates light-dependent seed germination. Plant Cell, 33(5), 1506–1529. https://doi.org/10.1093/plcell/koab060
Kozomara, A., Birgaoanu, M., & Griffiths-Jones, S. (2019). miRBase: From microRNA sequences to function. Nucleic Acids Research, 47(D1), D155–D162. https://doi.org/10.1093/nar/gky1141
Lanet, E., Delannoy, E., Sormani, R., Floris, M., Brodersen, P., Crété, P., Voinnet, O., & Robaglia, C. (2009). Biochemical evidence for translational repression by Arabidopsis microRNAs. Plant Cell, 21(6), 1762–1768. https://doi.org/10.1105/tpc.108.063412
Li, D., Wang, L., Liu, X., Cui, D., Chen, T., Zhang, H., Jiang, C., Xu, C., Li, P., Li, S., Zhao, L., & Chen, H. (2013). Deep sequencing of maize small RNAs reveals a diverse set of microRNA in dry and imbibed seeds. PLoS ONE, 8(1), e55107. https://doi.org/10.1371/journal.pone.0055107
Li, Z., Yang, J., Cai, X., Zeng, X., Zou, J.-J., & Xing, W. (2024). A systematic review on the role of miRNAs in plant response to stresses under the changing climatic conditions. Plant Stress, 14, 100674. https://doi.org/10.1016/j.stress.2024.100674
Lorenz, R., Bernhart, S. H., Höner zu Siederdissen, C., Tafer, H., Flamm, C., Stadler, P. F., & Hofacker, I. L. (2011). ViennaRNA Package 2.0. Algorithms for Molecular Biology, 6, 26. https://doi.org/10.1186/1748-7188-6-26
Luján-Soto, E., Juárez-González, V. T., Reyes, J. L., & Dinkova, T. D. (2021). MicroRNA Zma-miR528 versatile regulation on target mRNAs during maize somatic embryogenesis. International Journal of Molecular Sciences, 22(10), 5310. https://doi.org/10.3390/ijms22105310
Ma, C., Burd, S., & Lers, A. (2015). miR408 is involved in abiotic stress responses in Arabidopsis. The Plant Journal, 84(1), 169–187. https://doi.org/10.1111/tpj.12999
Martinez-Seidel, F., Beine-Golovchuk, O., Hsieh, Y.-C., & Kopka, J. (2020). Systematic Review of Plant Ribosome Heterogeneity and Specialization. Frontiers in Plant Science, 11. https://doi.org/10.3389/fpls.2020.00948
Motomura, K., Le, Q. T., Kumakura, N., Fukaya, T., Takeda, A., & Watanabe, Y. (2012). The role of decapping proteins in the miRNA accumulation in Arabidopsis thaliana. RNA Biology, 9(5), 644–652. https://doi.org/10.4161/rna.19877
Nieves-Cordones, M., Robles, P., & Quesada, V. (2025). Mutations in the plant-conserved uL1m mitochondrial ribosomal protein significantly affect development, growth and abiotic stress tolerance in Arabidopsis thaliana. Plant Growth Regulation, 105, 429–448. https://doi.org/10.1007/s10725-025-01282-x
Nonogaki, H. (2010). MicroRNA gene regulation cascades during early stages of plant development. Plant and Cell Physiology, 51(11), 1840–1846. https://doi.org/10.1093/pcp/pcq154
Palatnik, J. F., Allen, E., Wu, X., Schommer, C., Schwab, R., Carrington, J. C., & Weigel, D. (2003). Control of leaf morphogenesis by microRNAs. Nature, 425(6955), 257–263. https://doi.org/10.1038/nature01958
Pan, J., Huang, D., Guo, Z., Kuang, Z., Zhang, H., Xie, X., Ma, Z., Gao, S., Lerdau, M. T., Chu, C., & Li, L. (2018). Overexpression of microRNA408 enhances photosynthesis, growth, and seed yield in diverse plants. Journal of Integrative Plant Biology, 60(4), 323–340. https://doi.org/10.1111/jipb.12634
Reyes, J. L., & Chua, N. H. (2007). ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant Journal, 49(4), 592–606. https://doi.org/10.1111/j.1365-313X.2006.02980.x
Rincon-Guzmán, A., Beltrán-Peña, E., Ortíz-López, A., & Sánchez de Jiménez, E. (1998). Ribonucleoprotein particles of quiescent maize embryonic axes. Plant Molecular Biology, 38(3), 357–364. https://doi.org/10.1023/a:1006020121351
Sano, N., Rajjou, L., & North, H. M. (2020). Lost in translation: Physiological roles of stored mRNAs in seed germination. Plants, 9(3), 347. https://doi.org/10.3390/plants9030347
Sarkar Das, S., Yadav, S., Singh, A., Gautam, V., Sarkar, A. K., Prasad, M., Muthamilarasan, M., & Bhatt, D. (2018). Expression dynamics of miRNAs and their targets in seed germination conditions reveals miRNA–ta-siRNA crosstalk as regulator of seed germination. Scientific Reports, 8, 1233. https://doi.org/10.1038/s41598-017-18823-8
Schneider, C., Rasband, W., & Eliceiri, K. (2012). NIH Image to ImageJ: 25 years of image analysis. Nature Methods, 9, 671–675. https://doi.org/10.1038/nmeth.2089
Song, Z., Zhang, L., Wang, Y., Li, H., Li, S., Zhao, H., & Zhang, H. (2018). Constitutive expression of miR408 improves biomass and seed yield in Arabidopsis. Frontiers in Plant Science, 8, 2114. https://doi.org/10.3389/fpls.2017.02114
Tang, X., Bian, S., Tang, M., Lu, Q., Li, S., Liu, X., Tian, G., Nguyen, V., Edward, W., Wang, A., Rothstein, S., Chen, X., & Cui, Y. (2012). MicroRNA-mediated repression of the seed maturation program during vegetative development in Arabidopsis. PLoS Genetics, 8(2), e1003091. https://doi.org/10.1371/journal.pgen.1003091
Tian, T., Zheng, Y., & Bailey-Serres, J. (2025). Translational regulation of plant stress responses: Mechanisms, pathways, and applications in bioengineering. Annual Review of Phytopathology, 63, 117–146. https://doi.org/10.1146/annurev-phyto-121823-032335
Wang, L., Liu, H., Li, D., & Chen, H. (2011). Identification and characterization of maize microRNAs involved in the very early stage of seed germination. BMC Genomics, 12, 154. https://doi.org/10.1186/1471-2164-12-154
Wang, Q., Wan, J., Dang, K., Meng, S., Hu, D., Lin, Y., Qiu, X., Guo, Z., Fu, Z., Ding, D., & Tang, J. (2023). zma-miR159 targets ZmMYB74 and ZmMYB138 transcription factors to regulate grain size and weight in maize. Plant Physiology, 193(4), 2430–2441. https://doi.org/10.1093/plphys/kiad455
Waskom, M. L. (2021). seaborn: Statistical data visualization. Journal of Open Source Software, 6(60), 3021. https://doi.org/10.21105/joss.03021
Xiong, W., Lan, T., & Mo, B. (2021). Extraribosomal Functions of Cytosolic Ribosomal Proteins in Plants. Frontiers in Plant Science, 12. https://doi.org/10.3389/fpls.2021.607157
Zhang, L., Chia, J.-M., Kumari, S., Stein, J. C., Liu, Z., Narechania, A., Maher, C. A., Guill, K., McMullen, M. D., & Ware, D. (2009). A genome-wide characterization of microRNA genes in maize. PLoS Genetics, 5(11), e1000716. https://doi.org/10.1371/journal.pgen.1000716
Zhang, Y., Yu, Y., Wang, C., Li, Z., Liu, Q., Xu, J., Liao, J., Wang, X., Qu, L., Chen, F., Xin, P., Yan, C., Chu, J., Li, H., & Chen, Y. (2013). Overexpression of microRNA OsmiR397 improves rice yield by increasing grain size and promoting panicle branching. Nature Biotechnology, 31(9), 848–852. https://doi.org/10.1038/nbt.2630
Zhang, Z., Yu, J., Li, D., Liu, F., Zhou, X., Wang, T., Ling, Y., & Su, Z. (2010). PMRD: Plant microRNA database. Nucleic Acids Research, 38(Database issue), D806–D813. https://doi.org/10.1093/nar/gkp818
Zhao, Y., Li, X., & Wang, H. (2025). MicroRNAs in plants development and stress resistance. Plant, Cell & Environment, 48(4), 1–22. https://doi.org/10.1111/pce.15546
Downloads
Arquivos adicionais
Publicado
Como Citar
Edição
Seção
Licença
Copyright (c) 2026 Scientia Agropecuaria

Este trabalho está licenciado sob uma licença Creative Commons Attribution-NonCommercial 4.0 International License.
Los autores que publican en esta revista aceptan los siguientes términos:
a. Los autores conservan los derechos de autor y conceden a la revista el derecho publicación, simultáneamente licenciada bajo una licencia de Creative Commons que permite a otros compartir el trabajo, pero citando la publicación inicial en esta revista.
b. Los autores pueden celebrar acuerdos contractuales adicionales separados para la distribución no exclusiva de la versión publicada de la obra de la revista (por ejemplo, publicarla en un repositorio institucional o publicarla en un libro), pero citando la publicación inicial en esta revista.
c. Se permite y anima a los autores a publicar su trabajo en línea (por ejemplo, en repositorios institucionales o en su sitio web) antes y durante el proceso de presentación, ya que puede conducir a intercambios productivos, así como una mayor citación del trabajo publicado (ver efecto del acceso abierto).




